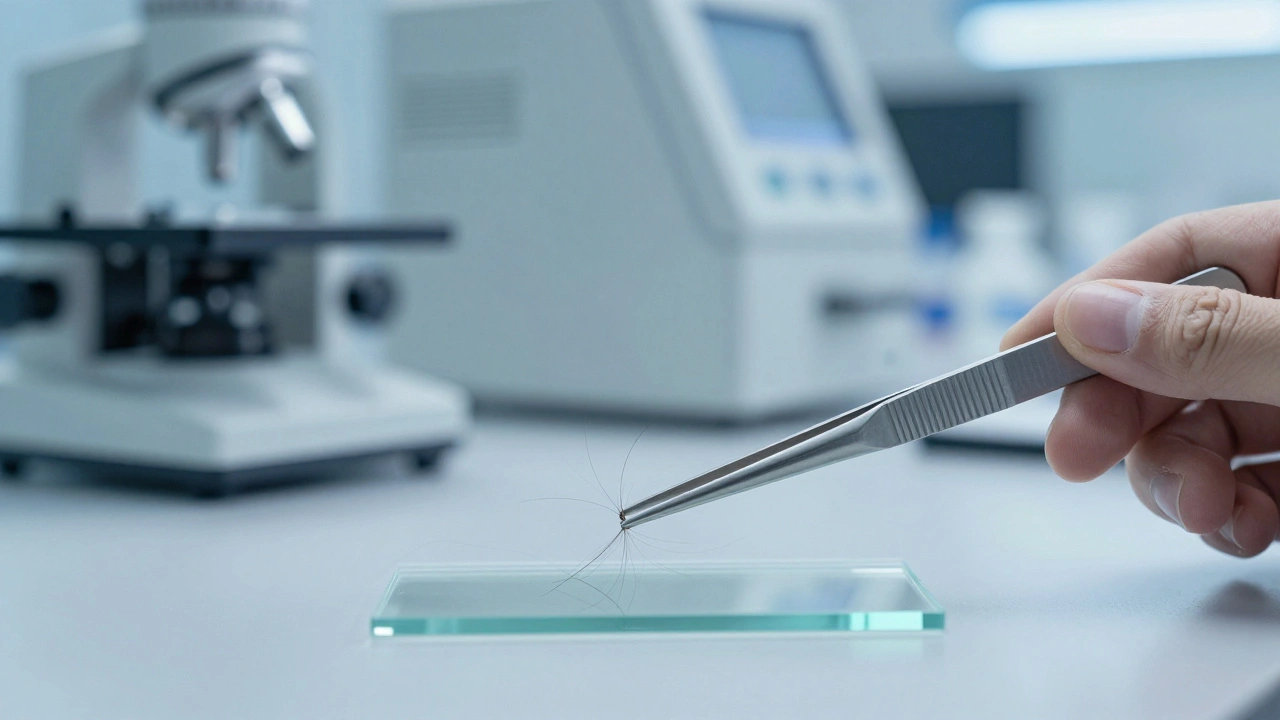
Imagine a forensic scientist analyzing a hair sample from a crime scene. They expect to find a clean, single genetic signature-a state called homoplasmy. But instead, they find two or more different genetic versions of mitochondrial DNA (mtDNA) within the same sample. This isn't a contaminated sample or a mixture of two different people; it's a biological phenomenon known as Mitochondrial DNA Heteroplasmy is the presence of multiple different mtDNA haplotypes within a single organism or cell. For a forensic investigator, this creates a massive headache: is this a rare genetic trait, a sequencing error, or a sign that the evidence came from multiple sources?
The Biological Reality of Heteroplasmy
To understand why heteroplasmy complicates forensic work, we first have to look at how mitochondria work. Unlike the DNA in your nucleus, which is a single pair of blueprints, each cell contains hundreds or thousands of mitochondria, and each mitochondrion has its own circular genome. In a perfect world, every single one of these circles would be identical. When they aren't, you have heteroplasmy.
This happens when a mutation occurs in some mitochondria but not others. This can happen at several levels: an entire person might have different mtDNA in different tissues, or a single cell might host a variety of different mtDNA molecules. In a forensic context, this means a swab from a cheek might show a different genetic profile than a sample from a hair follicle, even though they came from the same person. This lack of consistency can lead to "mismatches" during evidence comparison if the analyst isn't trained to recognize heteroplasmy.
The NUMT Problem: The Great Imposter
One of the biggest traps in forensic mtDNA analysis is the NUMT. A NUMT is a Nuclear Mitochondrial DNA segment-essentially a piece of mitochondrial DNA that jumped into the cell's nucleus. Because NUMTs look and act almost exactly like real mtDNA, sequencing machines often confuse the two.
Think of it like a photocopy of a document that accidentally got filed in the wrong folder. When a scientist runs a sequence, the machine might pick up a NUMT instead of actual mtDNA. This results in a "false positive" variant. For example, human chromosome 8 contains a nearly complete copy of the mtDNA sequence. If an analyst mistakes a NUMT for a mitochondrial mutation, they might report a heteroplasmy that doesn't actually exist, potentially leading to the wrong conclusion about a suspect's identity.
| Feature | True Heteroplasmy | NUMTs (Nuclear Segments) |
|---|---|---|
| Location | Mitochondrial Organelle | Cell Nucleus |
| Cause | Somatic or Germline Mutation | DNA Translocation/Insertion |
| Sequence Similarity | Identical to mtDNA backbone | ~86% average similarity to mtDNA |
| Forensic Impact | Intra-individual variation | False positive variant calls |
Technical Hurdles in Detection and Quantification
Detecting heteroplasmy isn't as simple as flipping a switch. It requires high-precision tools and a very specific approach to data analysis. The accuracy of the result depends heavily on how the DNA is "enriched"-basically, how the scientists isolate the mtDNA from the rest of the cellular junk.
Researchers often use a two-amplicon long-range PCR approach to clean up the sample. However, even with great isolation, the choice of the "reference sequence" (the gold standard map used for comparison) can change the result. If a scientist uses the rCRS (Revised Cambridge Reference Sequence) versus a combined rCRS and hg19 reference, they might see different results, especially for low-level variants. Specifically, any mutation appearing in less than 4% of the mitochondria is incredibly hard to distinguish from background noise or machine error.
There are also "danger zones" in the mtDNA genome. The D-loop region and the area between 5,000 and 10,000 base pairs are notorious for measurement artifacts. If a variant pops up here, a seasoned analyst will double-check it, knowing these regions are prone to errors that look like heteroplasmy but are actually just technical glitches.
Reporting Standards and Interpretation
Because every person has a little bit of genetic "noise," forensic labs can't report every single tiny difference. There is a standard threshold for reporting. Usually, if 20% or more of the mitochondria show a difference at a specific position, it's officially recorded as heteroplasmy. If it's lower than that, it might be dismissed as a sequencing artifact.
Reporting these values is a bit different from standard DNA testing. Normally, you'd just list the base pair (A, C, T, or G). But with heteroplasmy, you have multiple values at one spot. This requires a special notation system to show that a person has, for example, both an Adenine and a Guanine at the same position. This distinction is the difference between identifying a unique biological marker and misidentifying a sample due to contamination.
The Bigger Picture: Health and Evolution
While forensic scientists focus on identification, heteroplasmy has massive implications for medicine. It's the driving force behind many mitochondrial diseases. Because mtDNA doesn't follow Mendelian inheritance (you only get it from your mother), the level of heteroplasmy determines whether a person is healthy or suffers from a debilitating metabolic disorder. The "shift" in these levels-whether through random drift or natural selection-can determine if a mutation is purged from the population or passed on to the next generation.
Can heteroplasmy make a person's DNA look like it came from two different people?
Yes, in some cases. Because different tissues (like blood vs. hair) can have different levels of heteroplasmy, a forensic analyst might see different mtDNA profiles from the same individual. If they aren't aware of heteroplasmy, they might incorrectly conclude the samples came from two different people.
What is the difference between homoplasmy and heteroplasmy?
Homoplasmy is when every copy of mtDNA in a cell or organism is identical. Heteroplasmy is when there are two or more different versions (haplotypes) of mtDNA present.
Why are NUMTs such a problem for forensic labs?
NUMTs are sequences of mitochondrial DNA that have integrated into the nuclear genome. Because they look so similar to actual mtDNA, sequencing software often mistakes them for mitochondrial mutations, leading to false reports of heteroplasmy.
What is the typical reporting threshold for mtDNA heteroplasmy?
Most labs use a 20% threshold. This means that at least 20% of the sequenced mitochondria must carry the variant before it is officially reported as heteroplasmy rather than a technical error.
Does the choice of reference sequence affect the results?
Absolutely. The choice of reference (like rCRS or hg19) significantly impacts variant calling, especially for low-level heteroplasmy (below 4%), where the wrong reference can lead to incorrect identifications.
Next Steps for Analysts
If you're dealing with a suspected case of heteroplasmy, the first step is to verify the sample across multiple tissues if possible. If you only have one sample, focus on the D-loop and 5kb-10kb regions-if the variant is there, treat it with a high degree of skepticism until you can rule out a NUMT or an artifact.
For those moving toward Next-Generation Sequencing (NGS), the best move is to utilize a dedicated NUMT database. By filtering your results against known nuclear insertions, you can strip away the noise and find the true mitochondrial signal. Always document the enrichment strategy used, as this is the only way to ensure the results are reproducible and legally defensible in court.