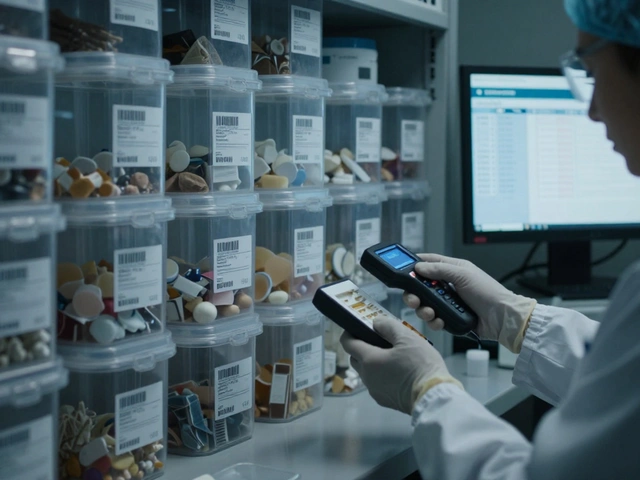
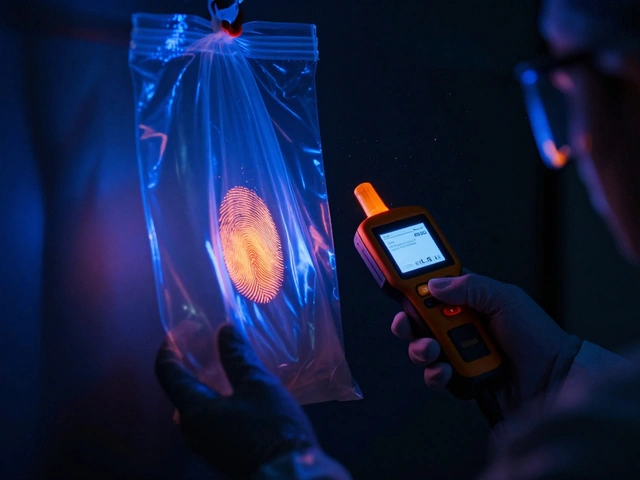
When people think of forensic DNA, they usually imagine a high-tech lab comparing the length of a few genetic segments to find a match. This is the traditional Short Tandem Repeat (STR) method. But what happens when the DNA is too degraded to work? Or when a sample is a messy mixture of three different people, and one person's DNA is barely there? This is where Non-STR DNA Markers step in. They aren't meant to replace STRs, but rather to fill the gaps when the standard toolkit fails.
Why SNPs Beat STRs in Tough Cases
If you're dealing with a cold case where the remains have been exposed to the elements for decades, STRs often fail. Why? Because STRs require relatively long, intact strands of DNA. SNPs only need a tiny fragment of DNA to be readable. Since they occur roughly every 300 to 500 base pairs, the chances of finding an intact SNP in a degraded sample are much higher.
Another huge difference is how they mutate. STRs change frequently, which is great for distinguishing between two cousins, but not for tracing deep history. SNPs are essentially "unique event polymorphisms" (UEPs). Once a mutation happens at a SNP site, it rarely happens again. This makes them incredibly reliable for building phylogenetic trees and determining deep ancestral lineages.
Mitochondrial DNA: The Powerhouse of Forensic Recovery
When a lab gets a sample consisting only of teeth, hair shafts, or old bones, they often turn to Mitochondrial DNA (or mtDNA). Unlike nuclear DNA, which you get from both parents, mtDNA is passed down only from the mother. This means every child of the same mother has the same mtDNA profile.
The real magic of mtDNA is its abundance. Every single cell has one nucleus but hundreds or thousands of mitochondria. If the nuclear DNA is completely gone, the mtDNA might still be there. Analysts typically focus on the hypervariable control region, which is packed with SNPs that can be sequenced to identify a maternal line. In extreme cases, if the control region is too common (like the most frequent Caucasian haplotype), scientists look into the coding regions for rarer SNPs to narrow things down.
Unlocking Paternal Secrets with Y-Chromosome Markers
In cases involving sexual assault or complex mixtures of male and female DNA, the Y-chromosome is the most valuable asset. Because the Y-chromosome is passed from father to son almost entirely intact, it allows investigators to isolate male DNA even when it's overwhelmed by female DNA from a victim.
Forensic teams use a few different tools here:
- Y-STRs: Great for recent family matches (within a few hundred years).
- Y-SNPs: Used to define haplogroups, which are ancestral groups that can trace a lineage back thousands of years.
- Alu Insertions: These act like biallelic markers and are fantastic for predicting where a person's ancestors came from geographically.
One tricky part about Y-chromosome testing is that different companies use different "trees" to name these groups. You might see one vendor call a group "R1b1a2" while another uses a different label. However, if you look at the specific SNP marker-like M269-that name remains the same across all platforms.
Solving the "Masking" Problem with SNP-STRs
One of the biggest headaches for a forensic analyst is a "two-donor mixture." Imagine a sample where one person's DNA is present in a huge amount, and another person's DNA is barely there. The dominant DNA often "masks" the partial DNA, making the second person invisible to standard STR tests.
This is where SNP-STRs come into play. These are compound markers that link a biallelic SNP with an STR polymorphism. By targeting a genomic region that is unique, these markers can cut through the noise. If the dominant donor doesn't have a specific allele that the partial donor does, the analyst can clearly see the second person's contribution, even in a heavily unbalanced mix.
| Marker Type | Best Use Case | Stability | Lineage Focus |
|---|---|---|---|
| STRs | Individual identification | Medium (mutates often) | Recent Family |
| SNPs | Degraded samples / Ancestry | Very High (stable) | Deep Ancestry |
| mtDNA | Bones, teeth, hair | High | Maternal Line |
| Y-Markers | Male-specific / Mixtures | Varies | Paternal Line |
| SNP-STRs | Unbalanced DNA mixtures | High | Individual / Mixed |
Predicting Appearance and Ancestry
Beyond just identifying a person, SNPs can actually tell us what that person looked like. This is a game-changer for "unidentified persons" cases. By analyzing specific SNPs, forensic anthropologists can predict eye color with surprising accuracy. It's not as simple as one gene; it's a complex interaction of several regions, but SNP analysis can decode this pattern.
Then there's ethno-geographic ancestry. By using a panel of about 70 autosomal SNPs known as Ancestry Informative Markers (AIMs), labs can estimate the percentage of a person's heritage from groups like Sub-Saharan African, East Asian, Native American, or Caucasian. This was famously used to help identify victims of the World Trade Center disaster, where the DNA was so fragmented that traditional methods were nearly useless.

The Modern Lab Workflow
Moving from STRs to SNPs isn't as simple as flipping a switch. It requires different hardware and software. Many labs now use a hybrid approach, combining both marker types to squeeze every bit of information out of a sample. Software like GeneMarker HID helps analysts manage this by providing mixture deconvolution and support for both SNP and STR panels.
We are also seeing a shift toward Mass Parallel Sequencing (MPS) and Whole Genome Sequencing (WGS). Instead of looking at 20 specific spots in the genome, these technologies allow scientists to read millions of base pairs at once. This makes it possible to detect rare variants and kinship links that were previously invisible.
Why are SNPs better for degraded DNA than STRs?
SNPs involve a change in just one single nucleotide, meaning the target sequence is much shorter than an STR. Because DNA breaks into small fragments as it degrades, it's much easier for a lab to find a tiny, intact SNP sequence than a longer STR sequence.
Can SNPs identify a specific person as uniquely as STRs?
A single SNP is biallelic (only two possible versions), so it has much less discriminating power than a multiallelic STR. However, by using a large panel of many different SNPs, labs can achieve a level of discrimination that matches or even exceeds traditional STR profiles.
What is the difference between Y-STRs and Y-SNPs?
Y-STRs mutate relatively quickly and are great for finding a father or brother in a recent timeframe (roughly 500 years). Y-SNPs are very stable and are used to define haplogroups, allowing researchers to trace a male lineage back thousands of years to a common ancestor.
How does mtDNA help in forensic cases?
mtDNA is found in the mitochondria, not the nucleus. Since there are thousands of copies of mtDNA per cell, it often survives when nuclear DNA has vanished. It's specifically useful for identifying remains from hair, bones, or teeth and tracing maternal lines.
What are SNP-STRs used for?
They are used primarily in mixed DNA samples where one person's DNA is much more abundant than the other's. They help prevent the "masking" effect, allowing the analyst to identify the minority contributor who would otherwise be missed by standard STR tests.
Next Steps for Investigators
If you're working with a sample that's coming back as "inconclusive" or "too degraded" via standard STR profiling, don't give up. Your next move depends on the sample type:
- For skeletal remains or old hair: Request mtDNA sequencing of the control region.
- For male-female mixtures: Shift to Y-STR and Y-SNP analysis to isolate the male profile.
- For severely fragmented DNA: Move to a high-density autosomal SNP panel for ancestry and identification.
- For complex mixtures with unbalanced donors: Explore the use of SNP-STRs to find the "hidden" contributor.